Centimorgans Chart
Centimorgans Chart - Classically, the linkage between two loci can be measured in centimorgans (cm), which represents the percent chance that these two loci will recombine an odd number of times. I was reading a paper titled "tetrad analysis in plants and fungi finds large differences in gene conversion rates but no gc bias" Haldane's map is an example that might be suitable for a. Linkage (physical linkage) is a measure of closeness on a chromosome. The answer hinges on a crucial distinction, which the difference between linkage and linkage disequilibrium. Is there an estimate that breaks down this total by chromosome (including the sex chromosomes)? Suppose that you have two genes located adjacent to each. The recombination frequency of two genes on the same chromosome is expressed in centimorgans (cm). The centimorgan is not a measure of the physical distance between gene loci, but a measure of the linkage between loci. Two genes of a flower, one controlling blue (b) versus white (b) petals and the other controlling round (r) versus oval (r) stamens, are linked and are 10 map units apart. Is there an estimate that breaks down this total by chromosome (including the sex chromosomes)? Classically, the linkage between two loci can be measured in centimorgans (cm), which represents the percent chance that these two loci will recombine an odd number of times. The recombination frequency of two genes on the same chromosome is expressed in centimorgans (cm). I was reading a paper titled "tetrad analysis in plants and fungi finds large differences in gene conversion rates but no gc bias" Suppose that you have two genes located adjacent to each. Haldane's map is an example that might be suitable for a. 1% recombination frequency is a distance of 1 centimorgan (cm) on the gene map. Yes, using centimorgans is fine if you know the conditional probability of recombination given centimorgans. Linkage (physical linkage) is a measure of closeness on a chromosome. Using the rough estimate of 7,000 centimorgans in a human. Haldane's map is an example that might be suitable for a. The answer hinges on a crucial distinction, which the difference between linkage and linkage disequilibrium. Using the rough estimate of 7,000 centimorgans in a human. Yes, using centimorgans is fine if you know the conditional probability of recombination given centimorgans. The recombination frequency of two genes on the same. The centimorgan is not a measure of the physical distance between gene loci, but a measure of the linkage between loci. The recombination frequency of two genes on the same chromosome is expressed in centimorgans (cm). 1% recombination frequency is a distance of 1 centimorgan (cm) on the gene map. Is there an estimate that breaks down this total by. Suppose that you have two genes located adjacent to each. Using the rough estimate of 7,000 centimorgans in a human. Two genes of a flower, one controlling blue (b) versus white (b) petals and the other controlling round (r) versus oval (r) stamens, are linked and are 10 map units apart. Why do genetic testing companies (ftdna,ancestrydna,23andme) express dna shared. Suppose that you have two genes located adjacent to each. Yes, using centimorgans is fine if you know the conditional probability of recombination given centimorgans. I was reading a paper titled "tetrad analysis in plants and fungi finds large differences in gene conversion rates but no gc bias" Classically, the linkage between two loci can be measured in centimorgans (cm),. Using the rough estimate of 7,000 centimorgans in a human. Is there an estimate that breaks down this total by chromosome (including the sex chromosomes)? Suppose that you have two genes located adjacent to each. Linkage (physical linkage) is a measure of closeness on a chromosome. In the human genome 1 centimorgan is approximately 10 6 base pairs, so the. I was reading a paper titled "tetrad analysis in plants and fungi finds large differences in gene conversion rates but no gc bias" The recombination frequency of two genes on the same chromosome is expressed in centimorgans (cm). Haldane's map is an example that might be suitable for a. Suppose that you have two genes located adjacent to each. Why. Why do genetic testing companies (ftdna,ancestrydna,23andme) express dna shared in centimorgans (cm) instead of in number of base pairs or in percent? Suppose that you have two genes located adjacent to each. Classically, the linkage between two loci can be measured in centimorgans (cm), which represents the percent chance that these two loci will recombine an odd number of times.. The answer hinges on a crucial distinction, which the difference between linkage and linkage disequilibrium. Classically, the linkage between two loci can be measured in centimorgans (cm), which represents the percent chance that these two loci will recombine an odd number of times. The centimorgan is not a measure of the physical distance between gene loci, but a measure of. Yes, using centimorgans is fine if you know the conditional probability of recombination given centimorgans. Suppose that you have two genes located adjacent to each. Linkage (physical linkage) is a measure of closeness on a chromosome. I was reading a paper titled "tetrad analysis in plants and fungi finds large differences in gene conversion rates but no gc bias" In. 1% recombination frequency is a distance of 1 centimorgan (cm) on the gene map. Haldane's map is an example that might be suitable for a. The answer hinges on a crucial distinction, which the difference between linkage and linkage disequilibrium. Two genes of a flower, one controlling blue (b) versus white (b) petals and the other controlling round (r) versus. Haldane's map is an example that might be suitable for a. Yes, using centimorgans is fine if you know the conditional probability of recombination given centimorgans. Two genes of a flower, one controlling blue (b) versus white (b) petals and the other controlling round (r) versus oval (r) stamens, are linked and are 10 map units apart. The answer hinges on a crucial distinction, which the difference between linkage and linkage disequilibrium. Is there an estimate that breaks down this total by chromosome (including the sex chromosomes)? 1% recombination frequency is a distance of 1 centimorgan (cm) on the gene map. The recombination frequency of two genes on the same chromosome is expressed in centimorgans (cm). Linkage (physical linkage) is a measure of closeness on a chromosome. In the human genome 1 centimorgan is approximately 10 6 base pairs, so the. The centimorgan is not a measure of the physical distance between gene loci, but a measure of the linkage between loci. Using the rough estimate of 7,000 centimorgans in a human. Classically, the linkage between two loci can be measured in centimorgans (cm), which represents the percent chance that these two loci will recombine an odd number of times.Need a DNA Chart? Who are You Made Of?
Chart Understanding Your DNA Results
ISOGG Wiki
Beginner's Guide to Shared Who are You Made Of?
Chart Understanding DNA Relationships Living DNA
Beginner's Guide to Shared Who are You Made Of?
DNA Numbers What they mean and how to use them
Chart Understanding Dna Relationships Liv vrogue.co
Need a DNA Chart? Who are You Made Of?
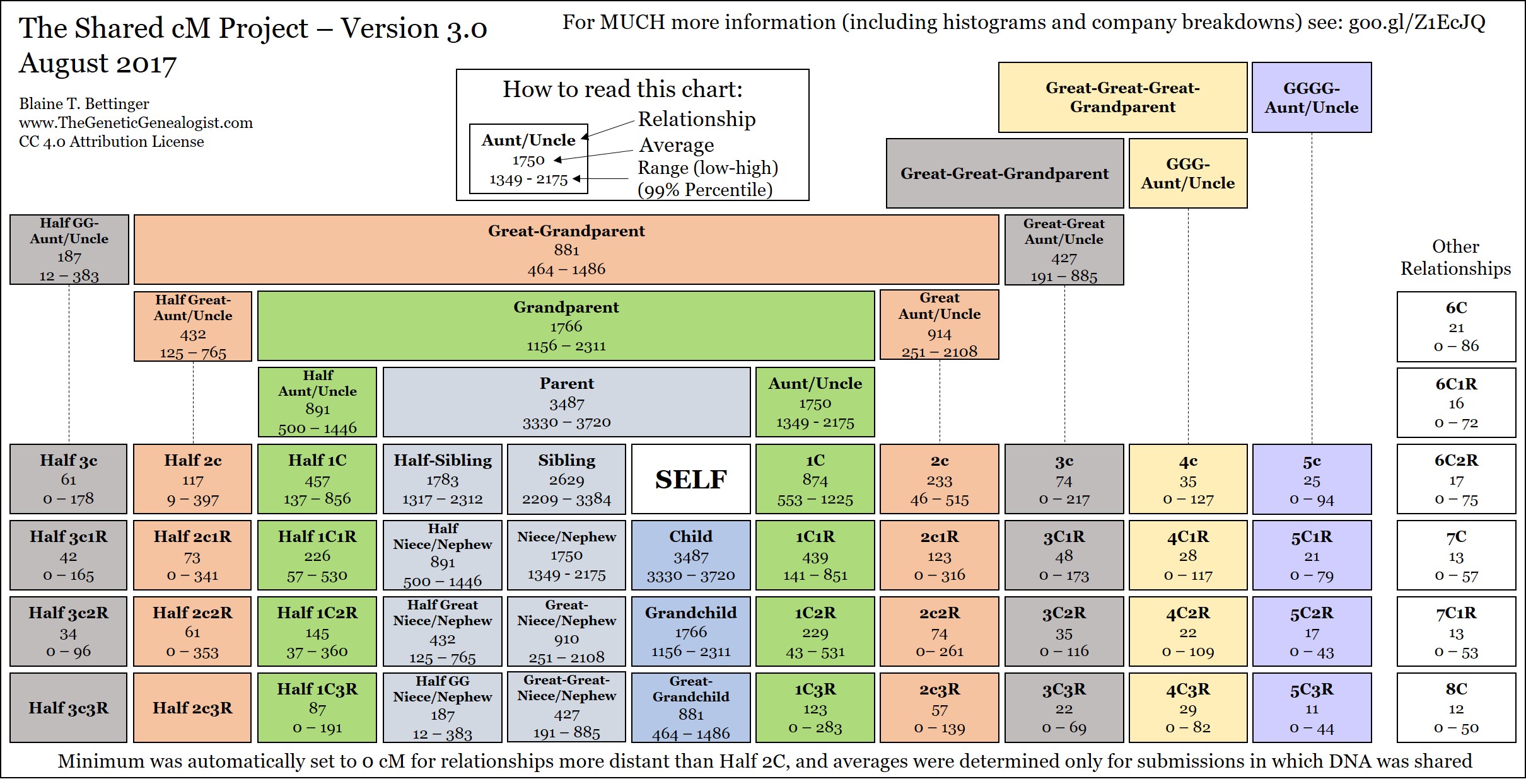
Compare for relationships. An interactive version of the shared cM data Dna
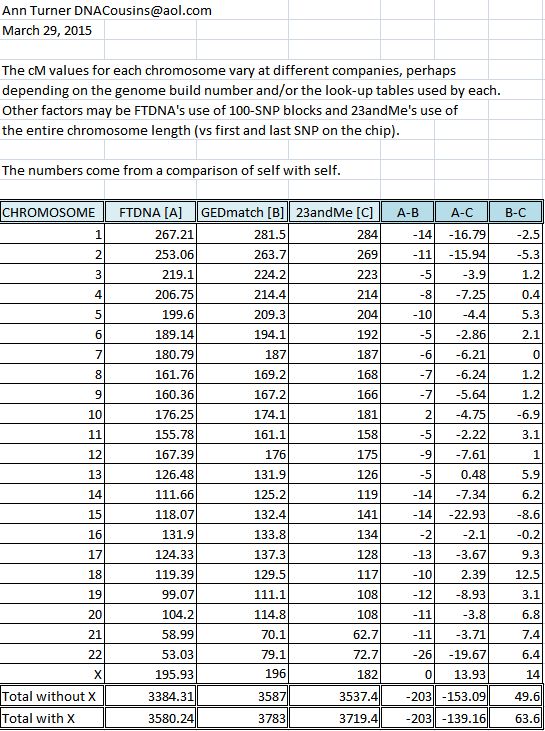
Why Do Genetic Testing Companies (Ftdna,Ancestrydna,23Andme) Express Dna Shared In Centimorgans (Cm) Instead Of In Number Of Base Pairs Or In Percent?
Suppose That You Have Two Genes Located Adjacent To Each.
I Was Reading A Paper Titled &Quot;Tetrad Analysis In Plants And Fungi Finds Large Differences In Gene Conversion Rates But No Gc Bias&Quot;
Related Post: